Documentation Index
Fetch the complete documentation index at: https://docs.nscale.com/llms.txt
Use this file to discover all available pages before exploring further.
Introduction
This cookbook walks you through creating a fine-tuning dataset from the PubMedQA corpus and running an end-to-end job on Nscale.
The goal is to demonstrate how to create a pair of CSV files in the required format (with columns question and answer) that you can upload to Nscale and use to fine-tune a base model.
Requirements
- Python 3.9+
- Packages: datasets, pandas
- Nscale service token and organization ID
Install dependencies:
pip install datasets pandas
Generate the dataset CSVs
To use Nscale to fine-tune a model, you need a dataset in the required CSV format. The script below converts a small subset of PubMedQA dataset (~2k samples) to a simple Q/A CSV format. You can adapt the same approach for other datasets.
- Input prompt: question
- Output target: long_answer when present, otherwise the categorical final_decision (e.g., yes/no/maybe)
It creates two CSVs: train and validation.
from typing import Dict, Iterable, Optional
import pandas as pd
from datasets import Dataset, DatasetDict, load_dataset
def example_to_qa_row(ex: Dict) -> Optional[Dict[str, str]]:
"""Convert one PubMedQA pqa_artificial example to a question/answer row.
- question: use the provided question string (omit context entirely).
- answer: prefer `long_answer` if non-empty; otherwise use `final_decision`.
"""
question = str(ex.get("question", "")).strip()
long_answer = ex.get("long_answer")
final_decision = ex.get("final_decision")
la = (str(long_answer).strip() if long_answer is not None else "")
fd = (str(final_decision).strip() if final_decision is not None else "")
answer = la if la else fd
return {"question": question, "answer": answer}
def dataset_to_csv(ds_split: Iterable[Dict], out_path: str) -> None:
rows = []
for ex in ds_split:
row = example_to_qa_row(ex)
if row:
rows.append(row)
pd.DataFrame(rows).to_csv(out_path, index=False)
def main() -> None:
"""Build train/validation Q/A split from sub-set of PubMedQA (pqa_artificial).
- Loads `qiaojin/PubMedQA` with config `pqa_artificial` (train split).
- Randomly splits into train and validation (seed=42), and select a subset of them.
- Select 2048 examples for training, and 128 for validation.
- Writes two CSVs with columns: `question`, `answer`.
- `answer` is `long_answer` if present, else `final_decision`.
"""
ds_all: DatasetDict = load_dataset("qiaojin/PubMedQA", "pqa_artificial")
if "train" not in ds_all:
raise RuntimeError("Expected a 'train' split in PubMedQA pqa_artificial")
base: Dataset = ds_all["train"]
split = base.train_test_split(test_size=0.2, seed=42, shuffle=True)
train_ds: Dataset = split["train"]
val_ds: Dataset = split["test"]
train_limit = min(len(train_ds), 2048)
val_limit = min(len(val_ds), 128)
train_head = train_ds.select(range(train_limit))
val_head = val_ds.select(range(val_limit))
dataset_to_csv(train_head, "pubmedqa_pqa_artificial_train_qa.csv")
dataset_to_csv(val_head, "pubmedqa_pqa_artificial_validation_qa.csv")
print(
"Wrote pubmedqa_pqa_artificial_train_qa.csv ({} examples) and "
"pubmedqa_pqa_artificial_validation_qa.csv ({} examples)".format(
train_limit, val_limit
)
)
if __name__ == "__main__":
main()
convert.py, then run it:
The script produces two files:
pubmedqa_pqa_artificial_train_qa.csvpubmedqa_pqa_artificial_validation_qa.csv
Each CSV has headers question,answer.
Create a fine-tuning job
You can create the fine-tuning job in the Nscale Console or via API. The steps below show how to do it with the Nscale Fine-tuning API.
1. Upload files
Export your Nscale token and organization ID, then upload both CSVs. Save the returned file ids for the next step.
export NSCALE_API_TOKEN="<your_service_token>"
export ORGANIZATION_ID="<your_organization_id>"
curl -X POST "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/files" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Content-Type: multipart/form-data' \
-H 'Accept: application/json' \
-F 'file=@pubmedqa_pqa_artificial_train_qa.csv'
curl -X POST "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/files" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Content-Type: multipart/form-data' \
-H 'Accept: application/json' \
-F 'file=@pubmedqa_pqa_artificial_validation_qa.csv'
2. Create the dataset
Use the two file ids returned above. Save returned dataset id to create a fine-tuning job:
curl -X POST "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/datasets" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Content-Type: application/json' \
-H 'Accept: application/json' \
-d '{
"name": "pubmedqa-pqa-labeled-qa",
"training_file_id": "<train_file_id>",
"validation_file_id": "<validation_file_id>"
}'
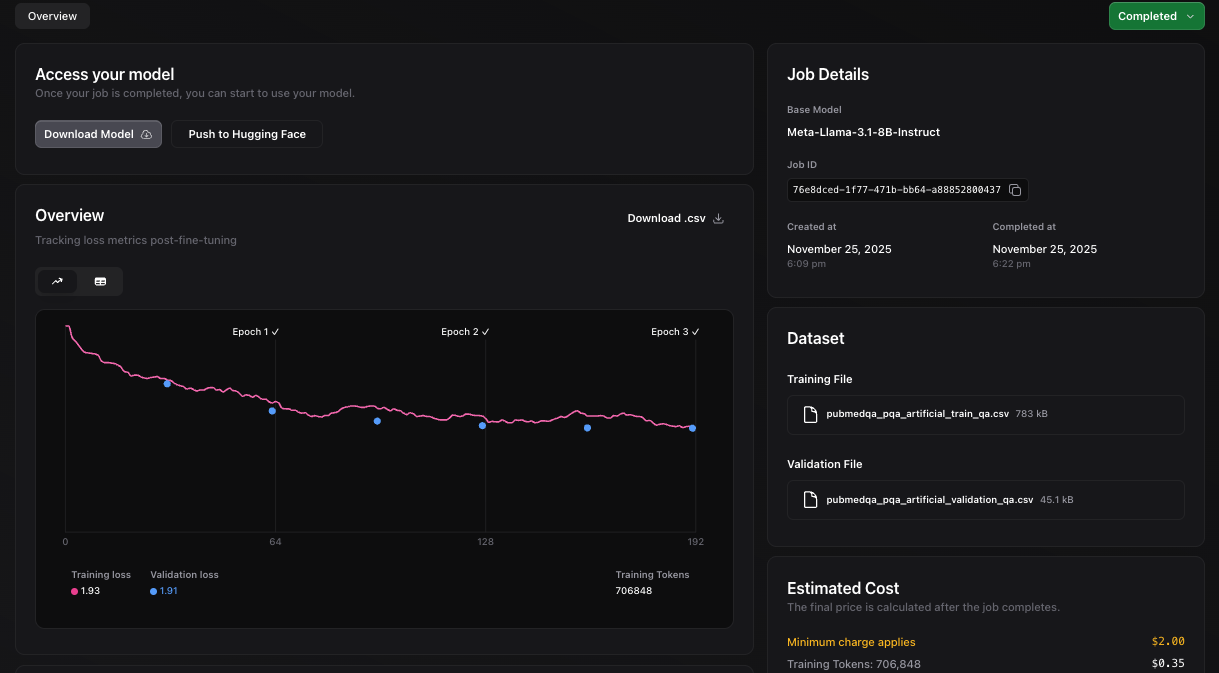
3. Create and monitor a fine-tuning job
Ensure you have enough credits in your account. First, list available base models and pick one. You can also find a list of available models in the Fine-tuning guide.
curl -X GET "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/base-models" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Accept: application/json'
question is the prompt and answer is the target output.
The example below uses meta-llama/Meta-Llama-3.1-8B-Instruct as the base model (ID: 5b912da4-4a68-43eb-9224-38239535d934). Replace base_model_id with the ID of your chosen model from the list above.
curl -X POST "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/jobs" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Content-Type: application/json' \
-H 'Accept: application/json' \
-d '{
"name": "pubmedqa-finetune",
"base_model_id": "5b912da4-4a68-43eb-9224-38239535d934",
"dataset": {
"id": "<YOUR_DATASET_ID>",
"prompt_column": "question",
"answer_column": "answer"
},
"hyperparameters": {
"batch_size": 32,
"best_checkpoints": false,
"learning_rate": 0.00001,
"lora": {
"alpha": 16,
"dropout": 0,
"enabled": true,
"r": 8,
"trainable_modules": []
},
"n_epochs": 3,
"n_evals": 6,
"warmup_ratio": 0,
"weight_decay": 0.03,
"mask_prompt_labels": false
}
}'
curl -X GET "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/jobs" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Accept: application/json'
curl -X GET "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/jobs/<job_id>" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Accept: application/json'
curl -X GET "https://fine-tuning.api.nscale.com/api/v1/organizations/$ORGANIZATION_ID/jobs/<job_id>/metrics" \
-H "Authorization: Bearer $NSCALE_API_TOKEN" \
-H 'Accept: application/json'